Our research focuses on the interactions between structure, mechanics, and function in both the plant and animal kingdoms. This will have a major impact on:
Plants
- developing designer cellulose fibers
- managing crops that are resistant to stresses and require fewer resources
Animals
- the ability to construct engineered tissues and organs
- creating new scaffolds for tissue repair and regeneration
- integrating implants in reconstructive surgery
- understanding the positive and negative effects of exercise, activity, and trauma
- providing therapies for tissue inflammation and fibrosis
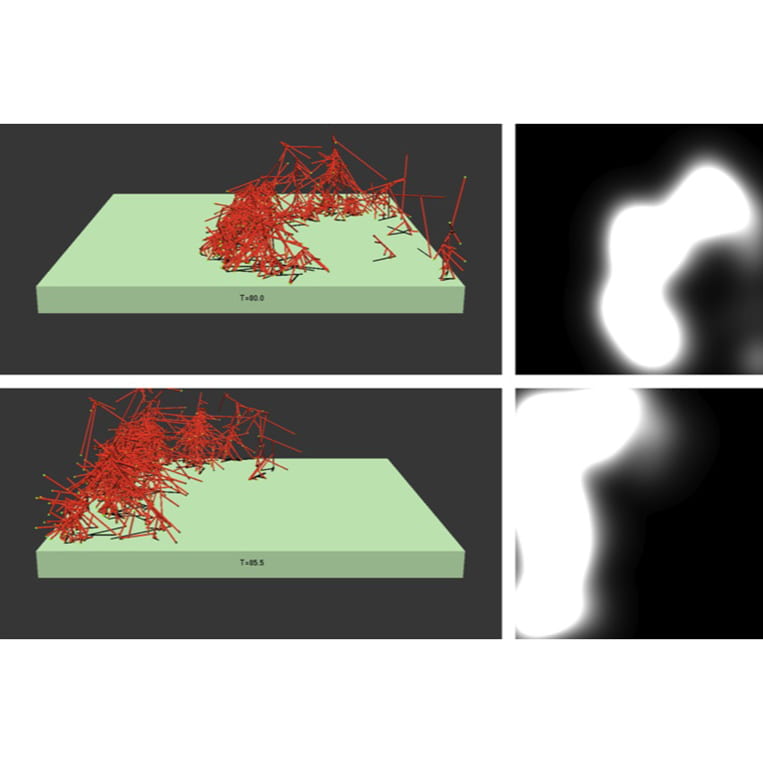

Carlsson Lab
Anders Carlsson’s work in biophysical modeling treats the molecular-scale processes that allow cells to move, divide and ingest nutrients from outside the cell.
Research Interests:
• Biophysical modeling
• Mechanobiology
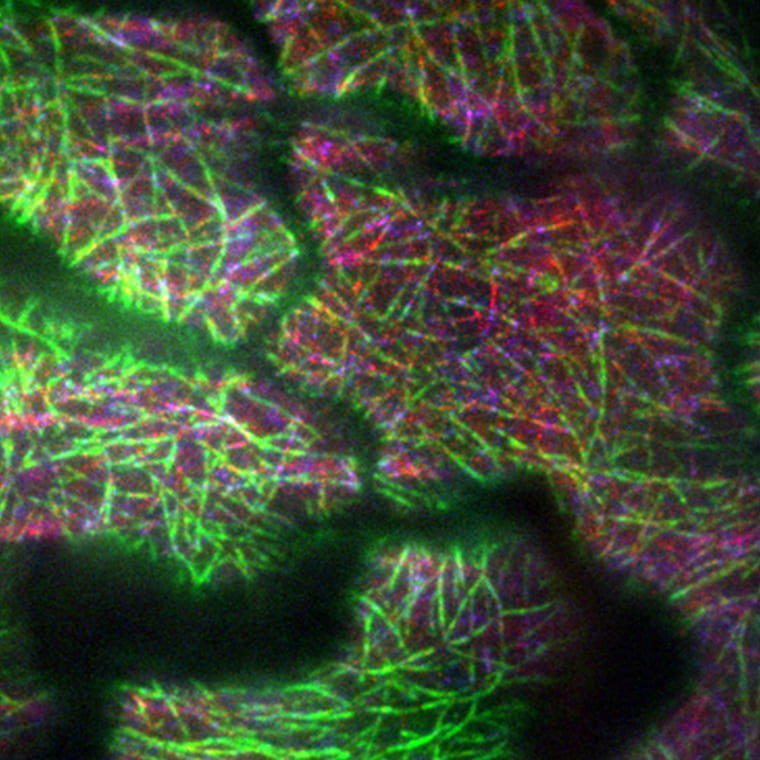
Dixit Lab
Ram Dixit’s research focuses on the cytoskeletal machinery for cell morphogenesis.
Research Interests:
• Mechanisms controlling microtubule dynamics and organization
• How cortical microtubules orchestrate cell wall deposition
• Role of mechanical stimuli in plant cell morphogenesis
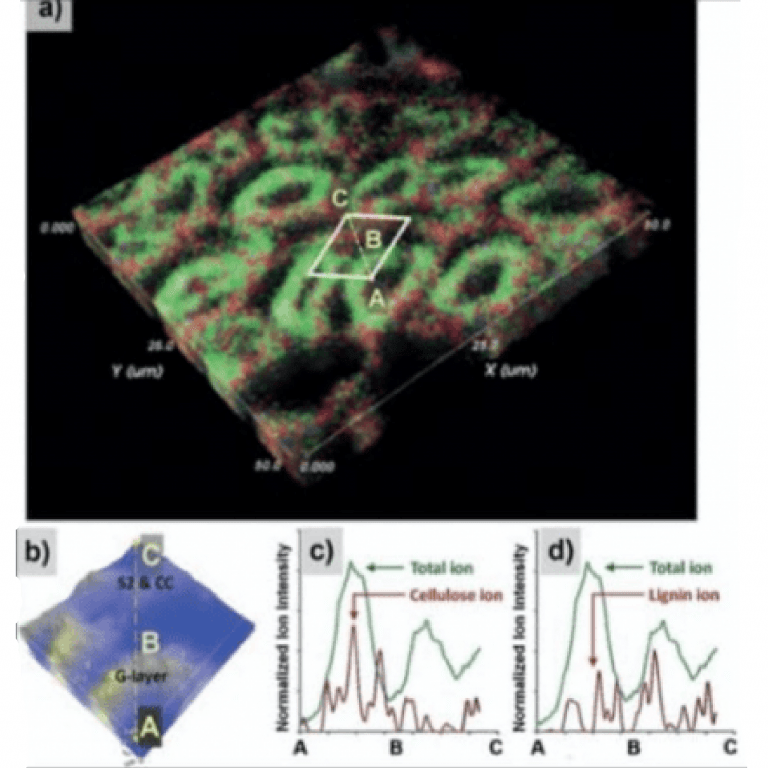
Foston Lab
Marcus Foston’s research seeks to develop innovative and novel routes to exploit and utilize lignocellulosic biomass, taking advantage of materials involved in industries such as agriculture, papermaking, and forestry products.
Research Interests:
• Sustainable conversion of biomass into chemicals using liquid-phase, heterogeneous catalysis.
• Interfacing the catalytic depolymerization of biomass with microbial utilization.
• Understanding how the plant cell respond to mechanical stimulus
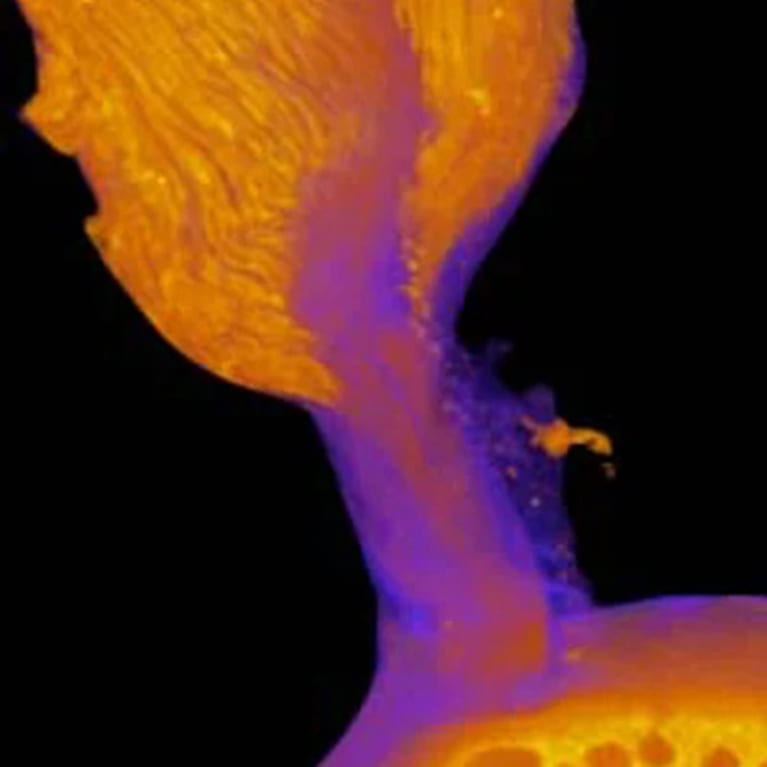
Genin Lab
Guy Genin’s research focuses on interfaces between tissues at the attachment of tendon to bone, between cells in cardiac fibrosis, and between protein structures at the periphery of plant and animal cells.
Research Interests:
• Mechanobiology
• Biomechanics
• Quantitative image analysis
• Interfaces and adhesion in nature, physiology, and engineering
Lab website
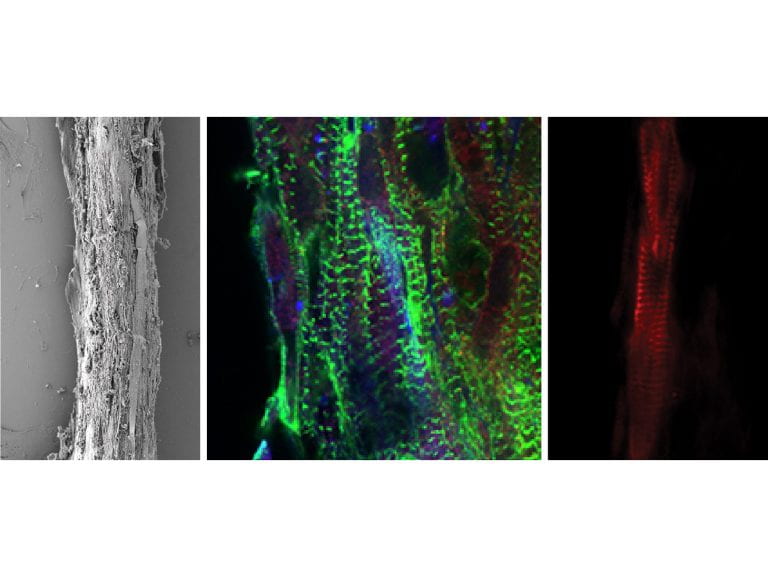
Huebsch Lab
Nathaniel Huebsch’s lab research focus is in basic and translational stem cell mechanobiology, with specific focus on hydrogels to control cell-mediated tissue repair, and 3-D iPSC-based heart-in-a-dish models to study the influence of mechanical loading and genetics on arrhythmia and contractility.
Research Interests:
• Pluripotent-stem cell derived cardiac micro-tissues for modeling cardiac development, drug toxicity and cardiomyopathy
• Synthetic Extracellular Matrix Mimetics with defined presentation of adhesion ligands and growth factors
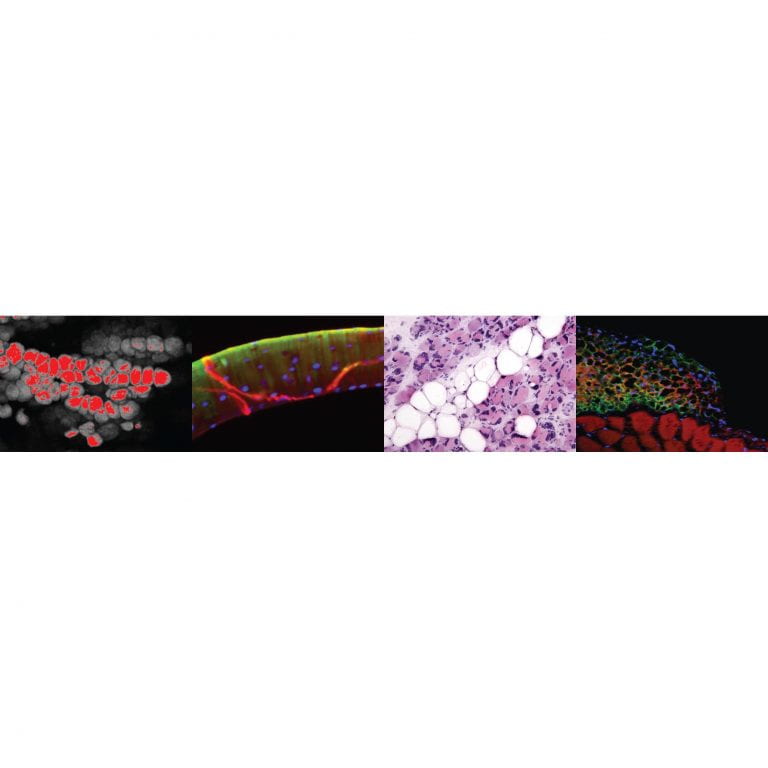

Meyer Lab
Gretchen Meyer’s lab focuses on understanding how changes at the cellular level in skeletal muscle affect muscle structure and function, and, in turn, how pathological changes in muscle affect cellular processes.
Research Interests:
• The functional consequences of fat accumulation in muscle
• The role of fat phenotype in fat-muscle cross-talk
• The cellular basis for poor muscle performance in diabetic peripheral neuropathy
• Guiding cell-based therapies to prevent fibrosis
Morley Lab
The Celeste Morley lab investigates how immune cells regulate their shape and migration patterns to combat pathogenic infections.
Research Interests:
• Identification of cells and genes required for resistance to pneumococcal infections.
• Structure-function analysis of the actin-bundling protein L-plastin – essential for T and B lymphocyte motility and activation.
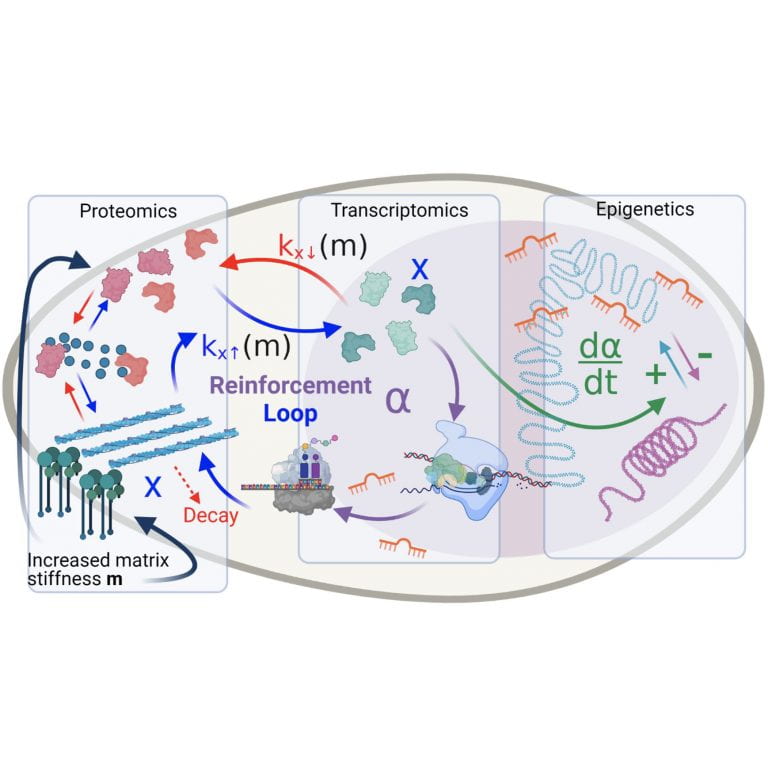
Pathak Lab
Amit Pathak’s lab studies cells in mechanically heterogeneous environments.
Research Interests:
• Regulation of tumor cell invasion by matrix properties
• Grouped cells in confined and heterogeneous environments
• Single cell migration in mechanically diverse environments
• Mechanical memory in cell migration
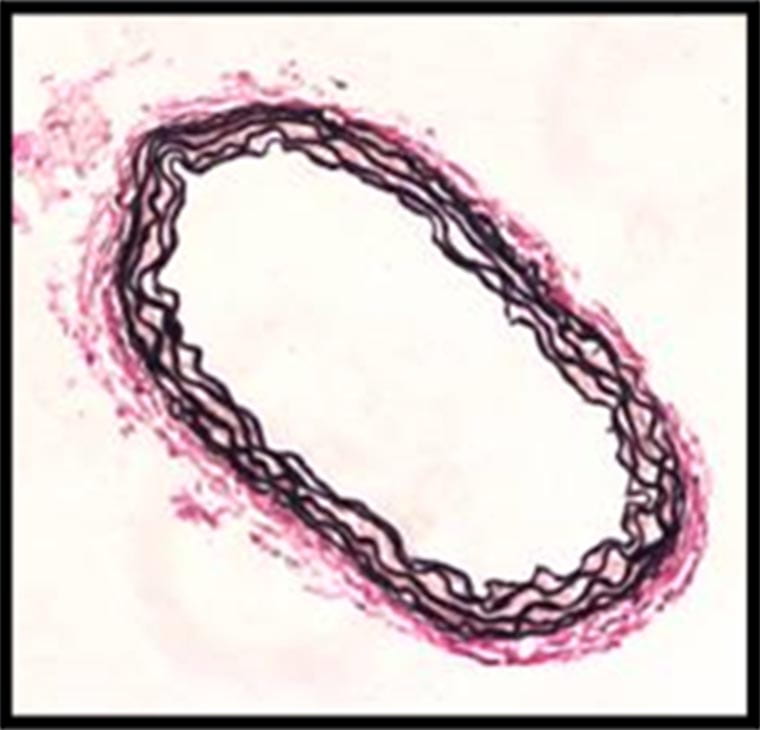
Wagenseil Lab
The Jessica Wagenseil lab studies vascular biomechanics and mechanobiology in development and disease. Her lab focuses on the mechanical behavior of large, elastic arteries during development and disease. They are interested in how the arterial wall remodels to maintain appropriate mechanical properties for heart function.
Research Interests:
• Cardiovascular mechanics, specifically in cardiovascular development
• Extracellular matrix proteins
• Microstructurally-based constitutive modeling